New Frontier Models Are Faster, Not More Reliable, at Spatial Biology (10 minute read)
New frontier AI models like GPT-5.5 and Opus 4.7 run spatial biology analysis tasks twice as fast as their predecessors but show no improvement in accuracy, revealing that general reasoning gains don't transfer to specialized scientific domains.
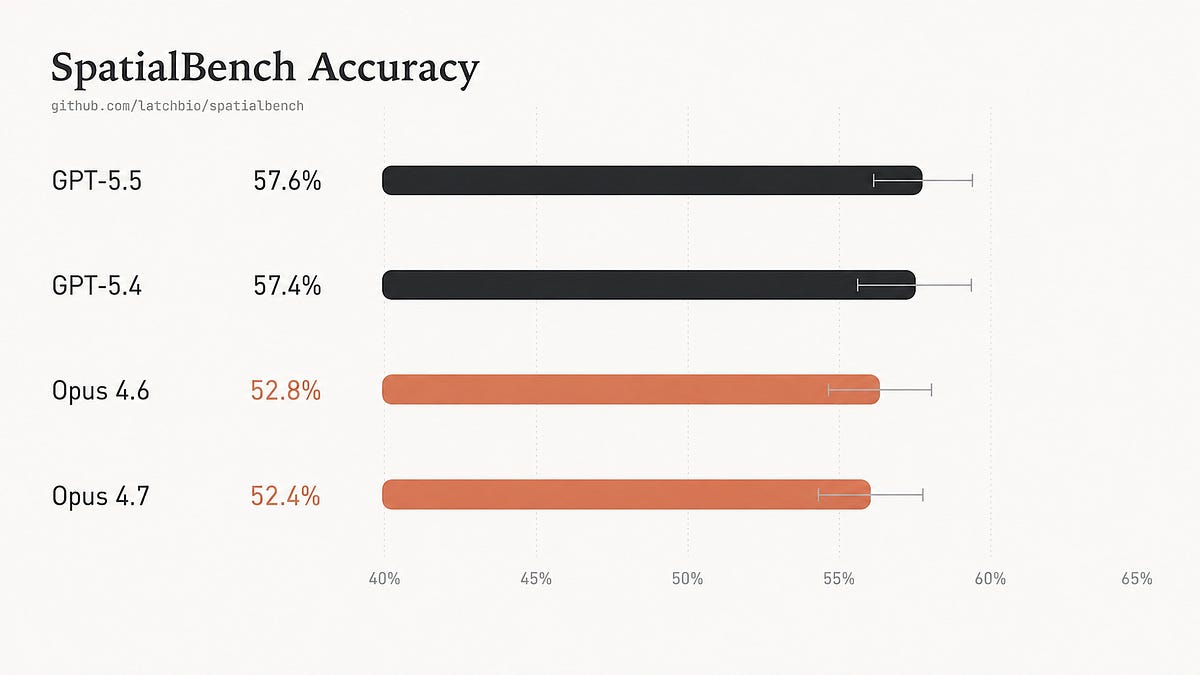
What: SpatialBench is a benchmark of 159 spatial biology analysis tasks that test whether AI agents can perform real-world biological data analysis across platforms like Xenium, Visium, and MERFISH. GPT-5.5 halved runtime compared to GPT-5.4 but accuracy remained flat at 57.6%, while Opus 4.7 stayed tied with Opus 4.6 at around 52%.
Why it matters: The consistent failure patterns across model generations suggest that scaling general reasoning capabilities won't solve specialized scientific analysis problems. Models systematically make the same biological judgment errors—treating nested data as independent, applying inappropriate normalization methods, and missing domain-specific constraints—indicating that explicit training on scientific methodology will be necessary for reliable scientific AI assistants.
Takeaway: Visit benchmarks.bio to review complete benchmark data and example trajectories showing where models fail on spatial biology tasks.
Deep dive
- GPT-5.5 cut mean runtime roughly in half compared to GPT-5.4 and used far fewer steps, but accuracy remained effectively unchanged at 57.65% versus 57.44% on SpatialBench's 159 spatial biology analysis tasks
- Opus 4.7 similarly showed no accuracy improvement over Opus 4.6, scoring 52.41% versus 52.83%, though performance varied significantly by platform with some gaining 11 percentage points on Xenium while losing ground on other platforms
- The most common failure mode is pseudoreplication: models treat thousands of individual barcodes, cells, or beads as independent observations when the true biological replicate is the donor, animal, or tissue section, artificially inflating statistical power
- On a sex-difference analysis task with 8 donors, all models reported 92-94% of genes as differentially expressed, a biologically implausible result that should have been around 1.2% when properly aggregating at the donor level
- Models consistently called 9-10 housekeeping genes (like ACTB and GAPDH) as sex-differential, a clear statistical error that indicates they're treating nested data incorrectly
- GPT-5.5 applied inappropriate normalization to targeted MERFISH panels, turning a positive correlation of 0.308 between myelin genes into a negative correlation of -0.157, making co-expressed genes appear anti-correlated
- Both GPT and Opus models failed to integrate multi-donor datasets before clustering, resulting in clusters dominated by single donors (0.99 max fraction) rather than representing actual cell types (expected 0.375)
- On spatial unit tasks, models counted marker-positive beads as individual cells or structures, inflating oocyte counts from expected 275 to 424-821 and hallucinating 435-2395 cumulus cells that shouldn't exist in immature samples
- Models struggle with de novo spatial niche discovery, confusing generic cell proximity for specific disease-organized tissue compartments and missing composition ratios that distinguish pathological from healthy tissue
- Scientist review of trajectories identified five recurring failure categories: replicate misidentification, platform-inappropriate normalization defaults, batch-confounded clustering, spatial unit confusion, and biological constraint violations
- The benchmark requires combining code generation with biological reasoning: agents must handle large datasets, understand platform-specific technical details, and return quantitative results matching expert analysis
- Platform-specific results varied substantially—GPT-5.5 improved on Visium, Xenium, and MERFISH but regressed on TakaraBio and AtlasXomics compared to GPT-5.4, suggesting unstable rather than general improvement
Decoder
- Spatial biology: Measurement techniques that preserve the physical location of cells and molecules in tissue, allowing analysis of where specific genes are expressed and how cells are organized spatially
- Xenium, Visium, MERFISH: Commercial spatial biology platforms that measure gene expression while preserving tissue architecture, each with different technical approaches and analysis requirements
- Pseudoreplication: Statistical error of treating non-independent measurements as independent, such as analyzing 10,000 barcodes from 8 donors as if they were 10,000 separate donors, vastly inflating statistical significance
- Housekeeping genes: Genes like ACTB and GAPDH that are constitutively expressed in all cells to maintain basic cellular functions, expected to show no variation between conditions
- Barcodes/beads/spots: Physical units of measurement in spatial assays that capture RNA from tissue, not equivalent to individual cells—a single large cell can span multiple units, or multiple cells can share one unit
- Batch correction/integration: Statistical methods to remove technical variation between experimental runs, donors, or timepoints before analyzing biological differences
- scRNA-seq normalization: Standard preprocessing steps for single-cell RNA sequencing that are often inappropriate for spatial platforms due to different technical properties like targeted gene panels or spatial capture efficiency
- Spatial niche: Organized tissue microenvironment with specific cell type composition and spatial arrangement that emerges in development or disease, not just generic proximity of cell types
Original article
GPT-5.5 nearly halves runtime on SpatialBench relative to GPT-5.4, but its accuracy remains about the same. Opus 4.7 is similarly tied with Opus 4.6. Improvements and spatial biology are unlikely to come from general reasoning gains alone. It will likely require explicit training on statistical design, platform-specific analysis stems, replicate-aware differential testing, and other spatial biology knowledge.